
Delighted to share our work on reverse engineering and testing global gene regulatory circuitry of the cranial neural crest. Congratulations @RuthWilliamsOx but also Ivan Ferreira, rest of Sauka-Spengler Lab and our collaborators E Repapi, S Taylor and Jim R. Hughes
cell.com/developmental-…

When Jim R. Hughes asked us (@FoldingGenome) to “break 3C” we thought he was crazy... we’ve ended up with NuTi Capture-C as a result 🥜 🥜 🥜.
biorxiv.org/content/10.110…

Exited to see DeepC our model for predicting 3D genome folding from DNA sequence now out in Nature Methods | nature.com/articles/s4159… (rdcu.be/b8pq3) Thanks to Jim R. Hughes, Gerton Lunter and all others who contributed.

Really excited to work on this project with Cecilia Lindgren and Jim R. Hughes! Wellcome wellcome.org/grant-funding/…

#Superenhancers function in an orientation dependent manner,challenging #enhancerbiology paradigm.
Hele Francis josephblayney Maria C Suciu, PhD Caz Harrold Jim R. Hughes Damien Downes Marieke Oudelaar Nature Genetics Hannah Long James Davies Gene Regulation MRC WIMM
biorxiv.org/content/10.110…



My paper is on bioRxiv - woohoo! 🎉 this has been a great team effort and nice to see some of my DPhil work out there. Jim R. Hughes Hughes Group


These two paper show how little we know about gene promoters still, first questions the widely prevalent need for
Pol II pausing (Jim R. Hughes): biorxiv.org/content/10.110…
and second:
Bivalent promoters (@Jothi_Lab): biorxiv.org/content/10.110…

Our paper is out in Nature Communications: Dynamics of the 4D genome during in vivo lineage specification and differentiation. Great teamwork with Rob Beagrie, Jim R. Hughes and Hughes Group. See thread below. #3Dgenome #generegulation nature.com/articles/s4146…

Our first keynote speaker on the second day of our symposium, Prof. Jim Hughes (Jim R. Hughes) from University of Oxford, is with us with his presentation titled “Understanding the ‘Dark Matter’ of the Human Genome”!
#RSGTurkey2021

Only ~2% of the human genome codes for proteins.
A Wellcome Discovery award for Jim R. Hughes jdavieslab and Cecilia Lindgren aims to deepen our understanding of the other 98% and its role in disease.
Big Data Institute MRC WIMM
Music: MNDLSS

#Technology behind Nucleome Therapeutics used to identify the gene linked to doubling the risk of death from #COVID ! Congratulations to our co-founders Jim R. Hughes & James Davies! The story is out in Nature Genetics nucleome.com/news-publicati…


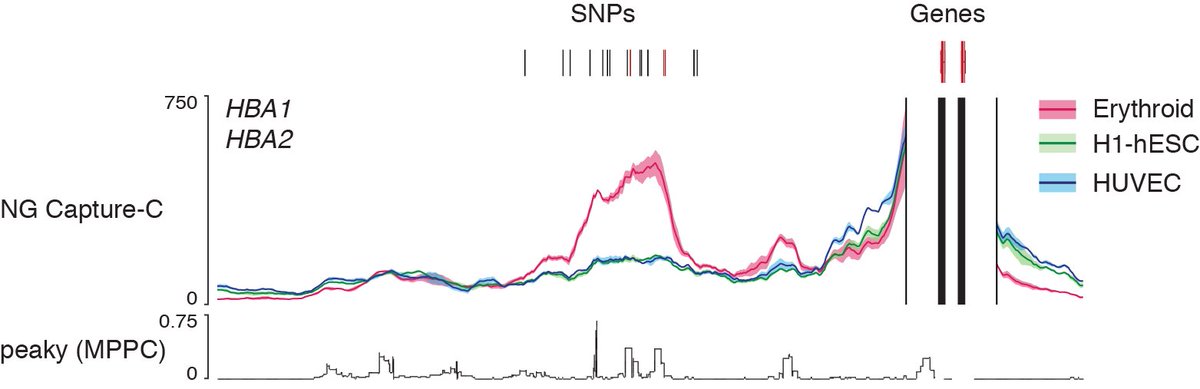
3) We use high-resolution NG Capture C (invented by James Davies and Jim R. Hughes) and statistical modelling to link SNPs to the genes they control.

How cis-regulatory elements interact to regulate gene expression-role of CTCF and cohesin- check out our pre-prints MRC WIMM Jim R. Hughes Hughes Group Rob Beagrie Jef Boeke Molecular Cell Nucleic Acids Res CTCF Papers 📚 Gene Regulation
biorxiv.org/content/10.110…
biorxiv.org/content/10.110…

Thank you Jim R. Hughes for the great lecture and for inspring MDC-BIMSB students! Updated list of upcoming lectures on '3D genomics' hosted by Prof Ana Pombo and Darío G. Lupiáñez attached.


It's going to be fun! We're looking forward to working with Cecilia Lindgren and the Big Data Institute over the next five years and beyond!
