Alexander Dietrich
@alexdietrich_
I am a PhD Student at the Chair of Experimental Bioinformatics at the Technical University of Munich
ID:1487022386067259392
28-01-2022 11:19:29
25 Tweets
49 Followers
83 Following


Check out our new paper on how bulk RNA-seq from blood offers insights into immune cell composition and specific immune responses to SARS-CoV-2 variants with just 10 million reads per sample. #RNASeq #ImmuneResponse
nature.com/articles/s4159… Markus List Jan Baumbach Alexander Dietrich

Check out the preprint for NeEDL (doi.org/10.1101/2023.1…), a method for identifying higher-order epistasis using PPI networks. Huge collaborative effort with Markus Hoffmann Nico Trummer Tim Kacprowski Jan Baumbach and many others.
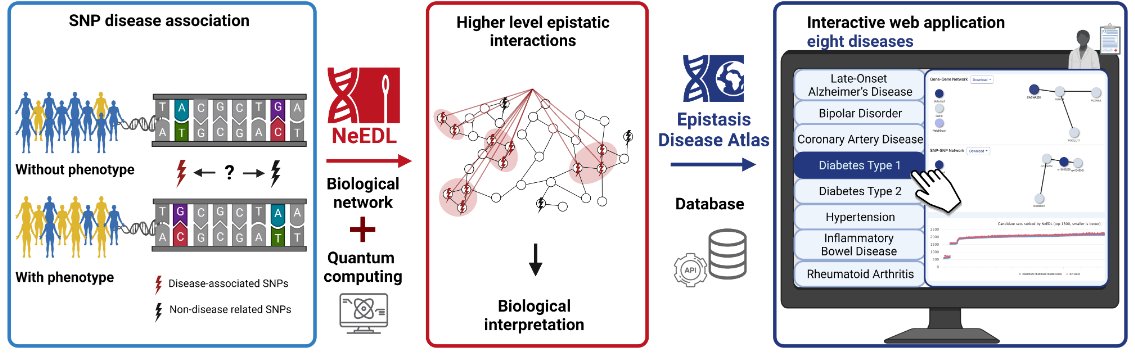

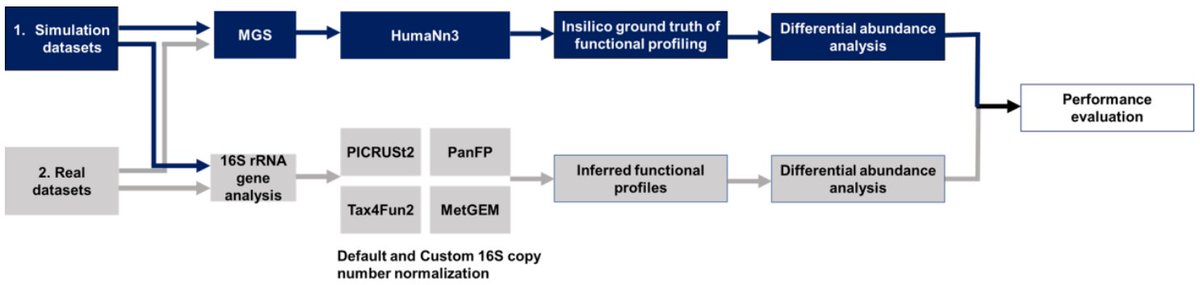
Looking for a weekend read? We benchmarked methods that infer functional profiles from 16S rRNA against matched metagenomes. These methods do not resolve health-related differences and should be used with caution. Great work by Monica Steffi Matchado w/ Tim Kacprowski Jan Baumbach et al.

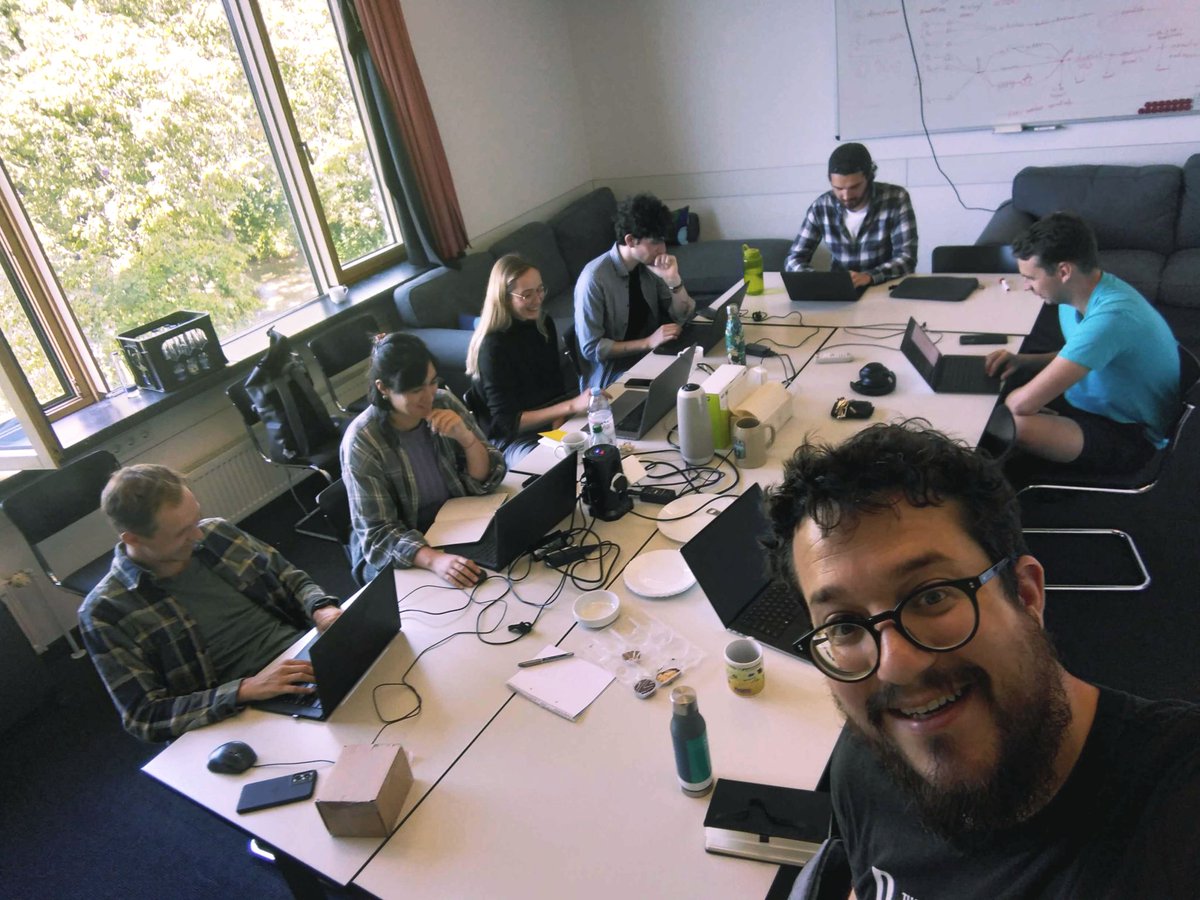
🌐 #Technicaluniversityofmunich
📅 Right now! 🙃
Short term scientific mission with Markus List and Florent Petitprez ‼️
🥇Figuring out the pipeline for creation of the very first myeloid cell atlas!
#Bioinformatics #BRCA COST
🖼️ Florent Petitprez #goodvibe


#ISMBECCB2023 ends today BUT there is still the Genetoberfest in Munich October 16-19 coming up. Submit an abstract before July 31st. Registration is free! Check out our great lineup of speakers genetoberfest.de Hope to see you there!


Alexander Dietrich will present omnideconv at poster C-109, a successor to the immunedeconv project focusing on second generation cell type deconvolution tools that learn celltype signatures from scRNA-seq data. Joint work with Francesca Finotello Lorenzo Merotto Gregor Sturm | [email protected] Federico Marini - @[email protected]


Hey #ismbeccb2023 if you are interested in pitfalls of machine learning and data leakage join us at 17:40 in the MLCSB session where Judith Bernett will talk about 'Cracking the black box of deep sequence-based protein-protein interaction prediction' doi.org/10.1101/2023.0…

📢 Our review on next-generation #deconvolution of the tumor microenvironment has been published on IRCMB! 🎉 shorturl.at/hlnX7
Kudos to Lorenzo Merotto Maria Zopoglou and Constantin Zackl!
Digital Science Center (DiSC) Universität Innsbruck @[email protected]


Our Genetoberfest TU München (16.-19.10.23, genetoberfest.de) will bridge bioinformatics and clinical research with exciting talks, panel discussions, flash talks, and posters. Registration is free of charge, submit an abstract easychair.org/conferences/?c… (deadline 17.07.23)


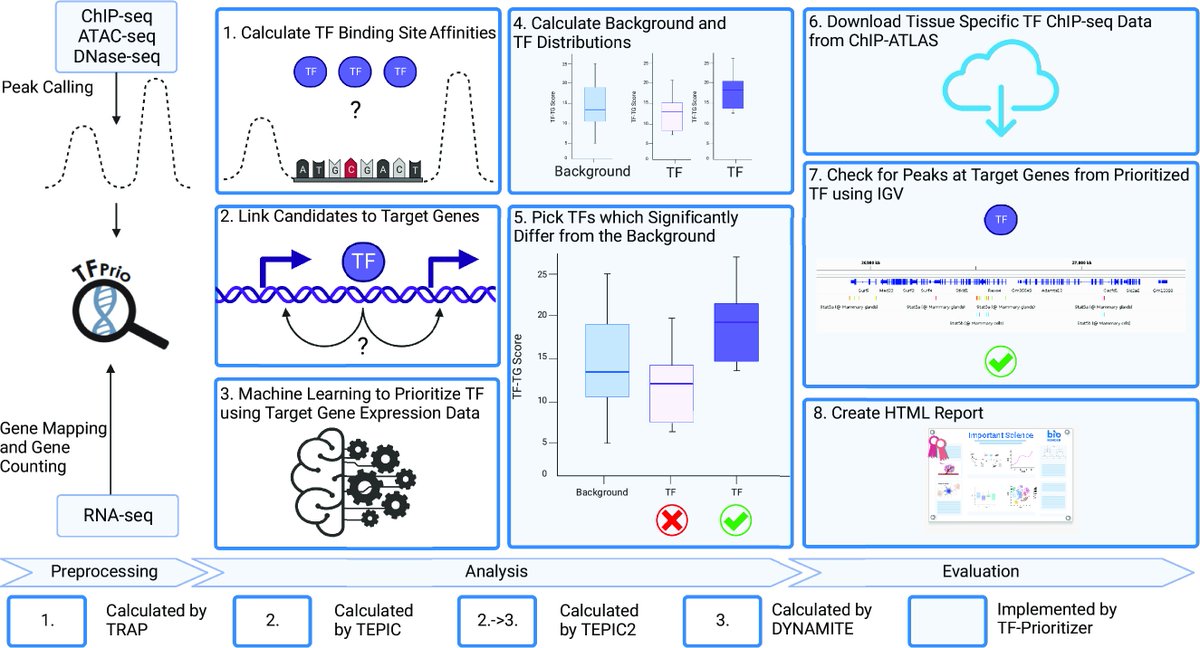
You have ChIP/ATAC- and RNA-seq data and want to learn more about essential transcription factors? Check out our tool TF-prioritizer which was just published in GigaScience with Markus Hoffmann Nico Trummer Marcel Schulz - @[email protected] CoSy.Bio David B. Blumenthal doi.org/10.1093/gigasc…

Our spongEffects method has now been published by OUP Bioinformatics doi.org/10.1093/bioinf… spongEffects extracts modules from ceRNA networks and calculates sample-specific enrichment scores, offering unique insights into miRNA regulation. Congrats to Fabio Boniolo Markus Hoffmann

📢 #PhD and #PostDoc positions in my group at Universität Innsbruck @[email protected] Digital Science Center (DiSC) to computationally reconstruct cancer interactomes from NGS multiomics
FWF project in collaboration with labi lab innsbruck Villunger Lab Joel Riley
euraxess.ec.europa.eu/jobs/75233
euraxess.ec.europa.eu/jobs/75236
Please RT! 🙏

📢4Y PostDoc position in my group at TU München to develop new comp. tools for the integration of single-cell RNA/ATAC-seq with spatial transcriptomics as part of the Novo Nordisk Foundation funded project MOPITAS w/ Oliver Stegle Richard Röttger kuba sędziński: careers.iscb.org/jobs/view/8391


Join us from Oct 16-20 TU München for the Genetoberfest! We seek to bridge (comp.) biomedical and clinical research and have a great lineup of speakers incl. Katie Hoadley Martin Hirst Michael Brudno 🎗️🇨🇦🇮🇱 Ana I Casas Kristiina Tammimies. Abstract submission is open at easychair.org/cfp/GO2023

Excited to introduce #circRNA sponging , a pipeline that systematically assesses the potential of circRNAs as biomarkers for diagnosis and treatment. Read our full article doi.org/10.1101/2023.0… #miRNA #sponging #circRNA #ceRNA Markus List Nico Trummer Jan Baumbach Katie Hoadley


Happy and excited to finally share this project!
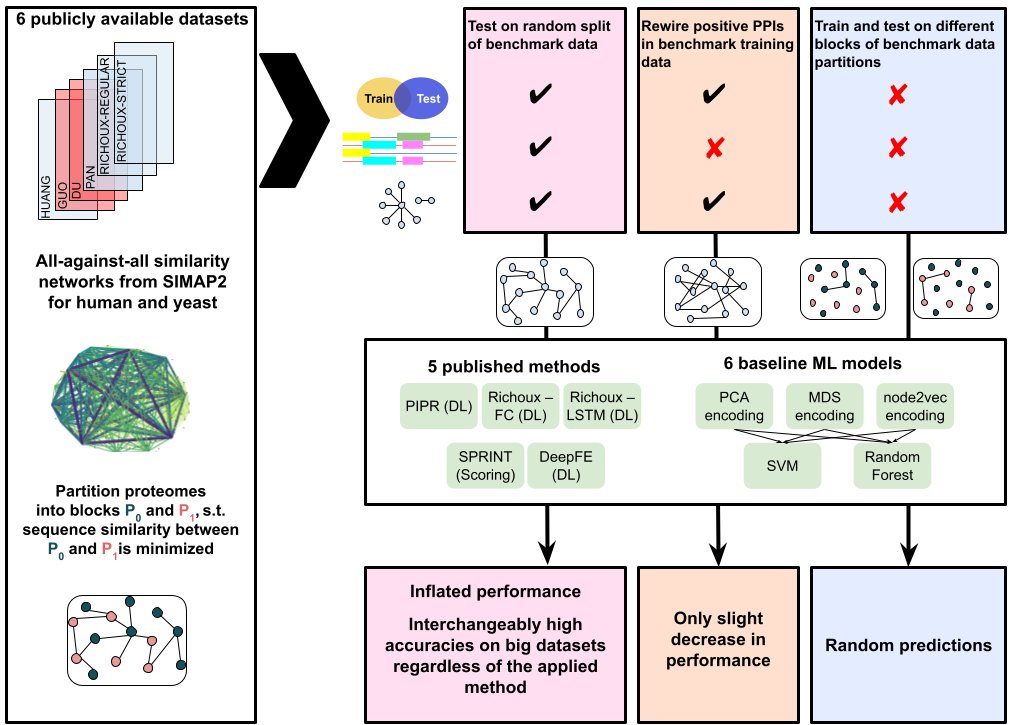
We show conclusively that high accuracies of deep learning-based PPI prediction models are exclusively due to data leakage via sequence similarities and node degree information.
biorxiv.org/content/10.110… Markus List David B. Blumenthal


📖Namco by Alexander Dietrich Monica Steffi Matchado Markus List et al. is a new web tool for comprehensive and user-friendly analysis of 16S data from raw processing down to classification, functional and network analysis.
Find out more in #MGen : doi.org/10.1099/mgen.0…
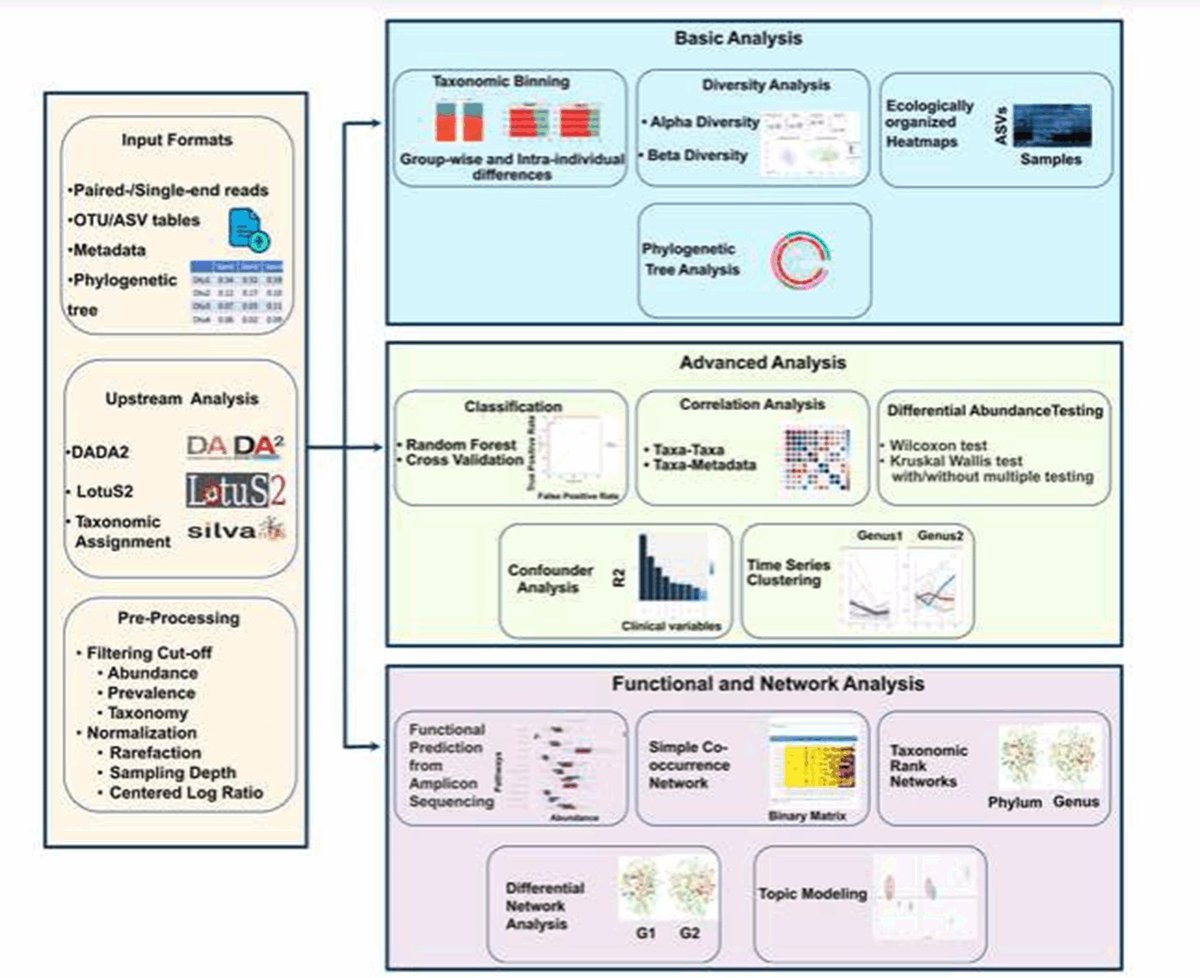

If you are at #ECCB2022 , pls come and visit poster “P018: Namco : the free microbiome explorer” to know more about our user friendly tool for microbial data analysis #microbiome Microbiome Signatures #microbialinformatics