
This #ReviewedPreprint makes use of AlphaFold2 to predict the models of tens of cohesin subcomplexes from different species. elifesciences.org/reviewed-prepr…
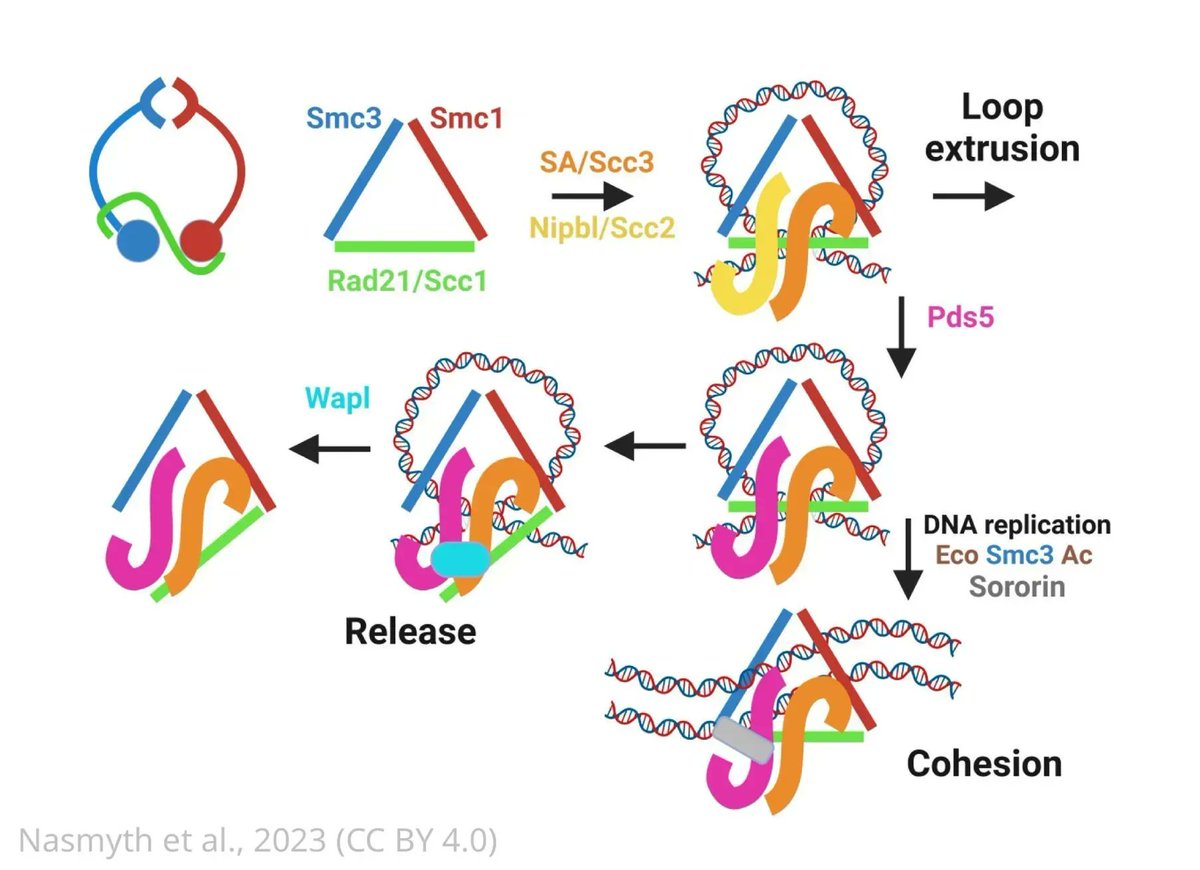


Antonio Chaves-Sanjuan Imaging_Artifact Have you tried Omokage search by PDBj?
pdbj.org/emnavi/omo-sea…
It only searches against experimental structures and not AlphaFold2 predicted structures though.


Check our latest work about AlphaFold2 and Deep Learning for Elucidating Enzyme Conformational Flexibility and Its Application for Design by Guillem Casadevall Cristina Duran
Silvia Osuna in JACS Au
iqcc.udg.edu/wordpress/2023…
#IQCCpaper #openaccess TCBioSys



#Alphafold2 on Colaboratory repeatedly disconnecting for no obvious reason after 5 hours on an A100 instance is not only super annoying but also quiet pricey! Anyone got a good step by step instruction for a local installation?

Fascinating read! RNA-guided endonucleases are present in all three domains of life! And happy to see how AlphaFold2 sheds light on the structure of SpuFz1 in complex with ωRNA-target DNA!! 🧬🔍 #ResearchBreakthrough #AlphaFold #CRISPR


today Alex TwentySeven(@univ_paris_cite,Inserm_EN,Université de La Réunion) is happy to present his work on 'An agnostic analysis of the human AlphaFold2 proteome using local protein conformations' at Jobim2023 (on the left results, on the right thinking Snoopy)






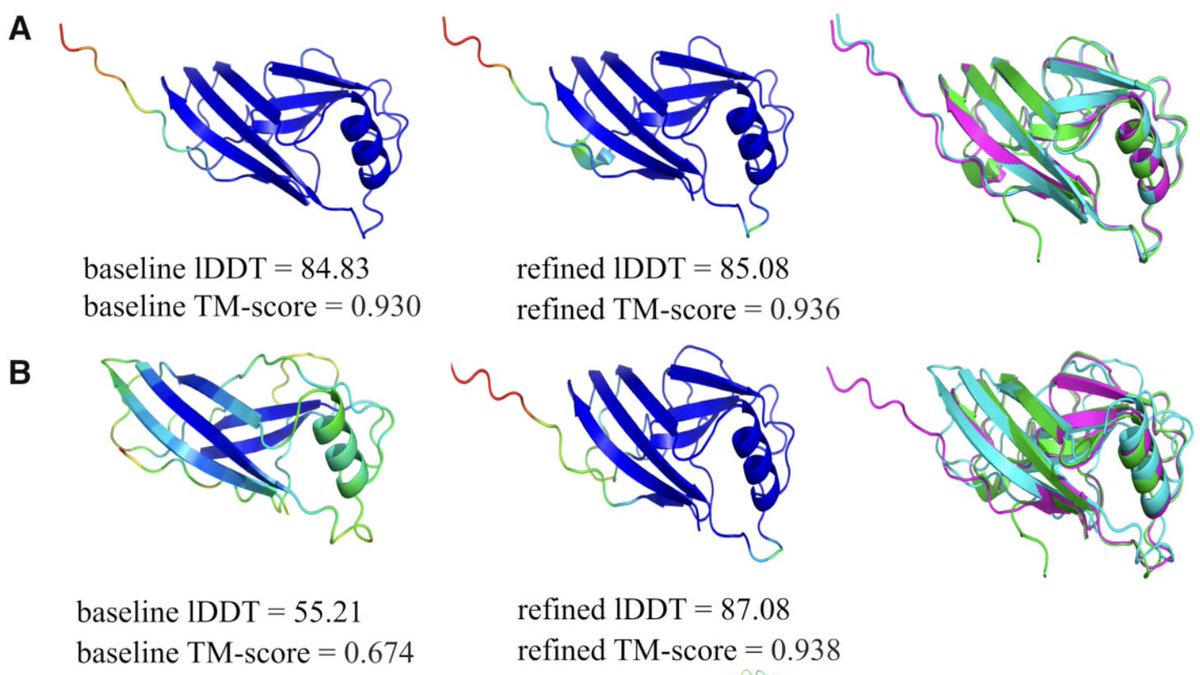
Improvement of protein tertiary and quaternary structure predictions using the ReFOLD refinement method and the AlphaFold2 recycling process Recep Adiyaman, PhD, Nicholas Edmunds, Ahmet G Genc, Shuaa M A Alharbi, Prof. Liam McGuffin in Bioinformatics Advances
doi.org/10.1093/bioadv…




Interested in #AlphaFold ?
Don’t miss Ron Vale’s #LMBSeminar on Monday 17th July at 4PM (BST). Ron is Executive Director at HHMI | Janelia and will be discussing ‘AlphaFold2 as a Tool for Cell Biology.’
More details: www2.mrc-lmb.cam.ac.uk/news-and-event…







