Benedict Paten
@BenedictPaten
Associate Professor at UCSC; Director, Computational Genomics Lab, UC Santa Cruz Genomics Institute
ID:716641980969684992
https://cgl.genomics.ucsc.edu/ 03-04-2016 15:02:17
517 Tweets
2,6K Followers
417 Following


Last chance to send your abstract to the International Genome Graph Symposium 2024 in Ascona! More info at iggsy.org
Keynotes include Arang Rhie Ben Langmead Camille Marchet @[email protected] Eimear Kenny Ewan Birney Jasmijn Baaijens Kateryna Marinka Zitnik Zamin Iqbal #iggsy24


Our CARD pilot paper with Kimberley Billingsley Cornelis Blauwendraat Benedict Paten and many others is online! We establish a wet + dry lab protocol for single nanopore flow cell sequencing, which generates state-of-the-art SNP and SV calls + assemblies. rdcu.be/dl901

A new set of Oxford Nanopore sequencing protocols, implemented in an Alzheimer's research project, will open up previously inaccessible regions of the genome with medically relevant variations. This is a huge step forward for large-scale genomics projects! 🎉
ucscgenomics.sites.ucsc.edu/2023/09/14/new…


Excited to share our collaboration with Dean lab National Cancer Institute! With long reads we sequenced a panel of cervical + head and neck cancer cell lines with complex SVs and aneuploidy. In particular, breakage-fusion-bridge (BFB) amplifications were ubiquitous (1/n) medrxiv.org/content/10.110…



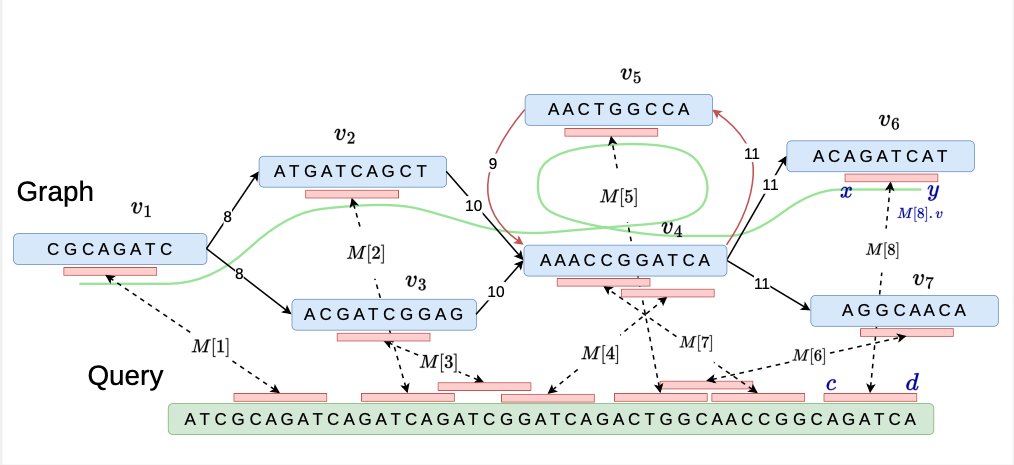
In a new paper with Jyotshna Rajput and Ghanshyam Chandra, we developed new algorithms for co-linear chaining on cyclic pangenome graphs. To appear in WABI
biorxiv.org/content/10.110…
github.com/at-cg/PanAlign…

A group of us have been thinking about how to explore human pangenome scale data and make them easier to understand for while. Great collaboration with Genome in a Bottle Fritz Sedlazeck Asif Khalak Sairam Behera.
twitter.com/naturemethods/…

Preprint on hifiasm ultra-long (UL) integration for better assembly. Now hifiasm works with
HiFi
HiFi + UL
HiFi + trio/Hi-C
HiFi + UL + trio/Hi-C
One executable and mostly one command line. Work by Haoyu Cheng, Mobin Asri, Julian Lucas, Sergey Koren. Help from HPRC and T2T.

Our latest methods paper 'Minmers are a generalization of minimizers that enable unbiased local Jaccard estimation' (aka MashMap3) is out with Bryce Kille Erik Garrison Todd J Treangen! There's an interesting backstory here 🧵 \ biorxiv.org/content/10.110…


As #PublicServiceRecognitionWeek comes to a close, I want to shout out my fellow Terp and Maryland resident, Adam Phillippy.
He is one of 13 Maryland finalists for The Partnership's Samuel J. Heyman Service to America Medals -- the sort of Oscars for public service workers.


The Human Pangenome Reference Consortium - the 3rd chapter. For an intro into the human pangenome, check out this thread - twitter.com/ewanbirney/sta… - and additional papers - twitter.com/ewanbirney/sta… but now on to the weirdest 'our genome does that!' moment. Brace yourself for genome geekery


We're ✨revolutionizing✨the human genome reference. The #HumanPangenome Consortium, co-led by UC Santa Cruz, has just published a draft of a more robust, diverse, and complete reference. This will be a game-changing resource for genomics & health research news.ucsc.edu/2023/05/pangen…

Researchers from the National Human Genome Research Institute-funded Human Pangenome Reference Consortium (@HumanPangenome) have completed a collection of new human reference genome sequences that much more accurately reflect global diversity! genome.gov/pangenome


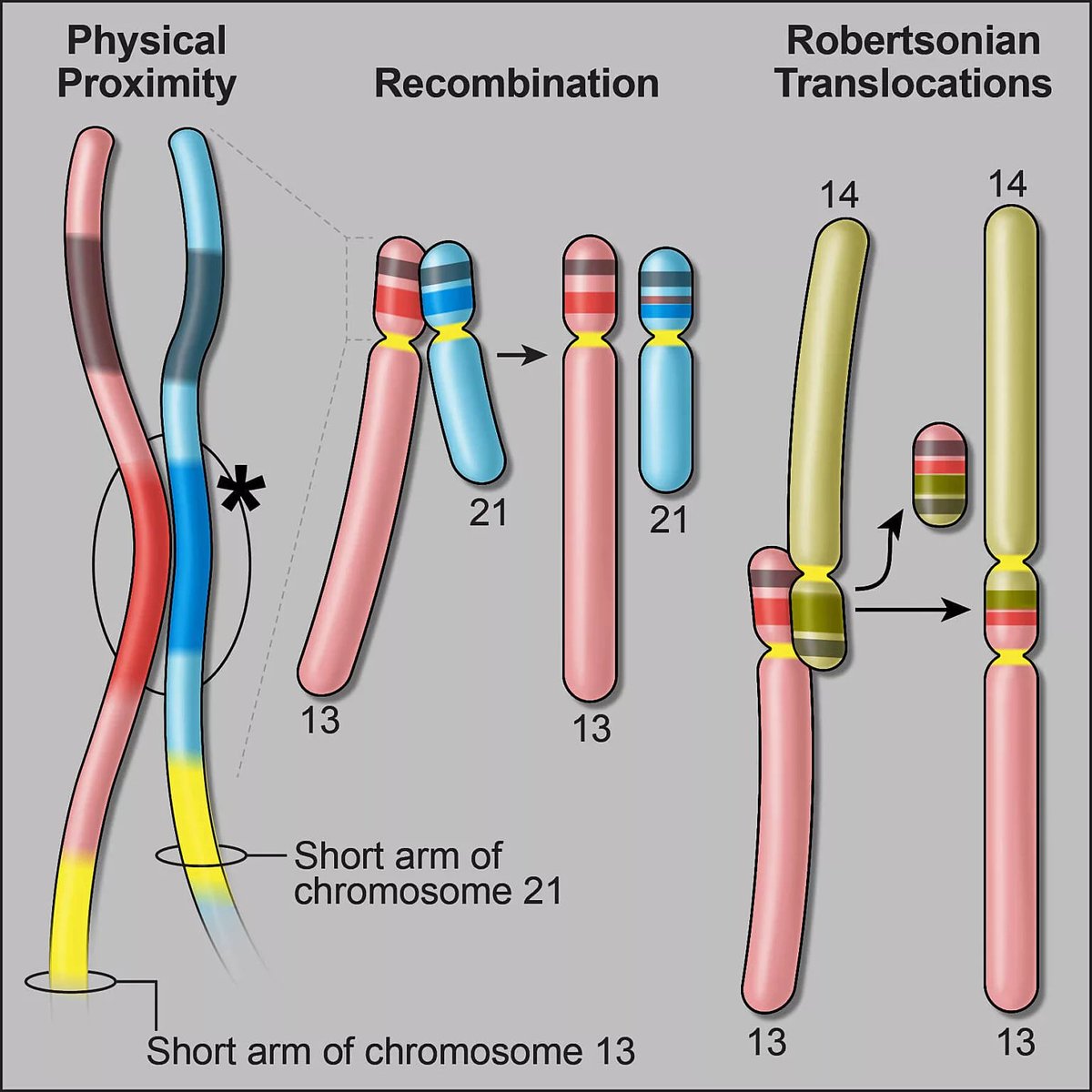
My favorite T2T/Pangenome discovery so far: a multi-megabase inverted segmental duplication on the short arm of chr14 that explains the mechanism of Robertsonian translocations. Great writeup by Stowers Institute 'Answering a 50-year-old mystery' stowers.org/news/investiga…

Congratulations to the Human Pangenome Reference Consortium - in particular Benedict Paten for his leadership + highlighting Ensembl's role via Fergal Martin in annotation. For the rest of this thread I will explain this technical genomics tour de force using... Shakespeare nature.com/articles/s4158…