
Haoyu Cheng
@ChengChhy
ID:1052555407275700224
17-10-2018 13:42:19
50 Tweets
267 Followers
227 Following



The previous hifiasm paper described haplotype-resolved assembly with trio data. In this new paper, Haoyu Cheng achieves chromosome-long phased assembly without parents. Tested for human and non-human species. A single executable and one command line for the entire process.



Hifiasm published in Nature Methods.
Preprint: arxiv.org/abs/2008.01237
Source code: github.com/chhylp123/hifi…
Assemblies: zenodo.org/record/4393631 & zenodo.org/record/4393750
Journal paper: nature.com/articles/s4159…
Fine work by Haoyu Cheng gconcepcion.bsky.social Xiaowen Feng Haowen Zhang

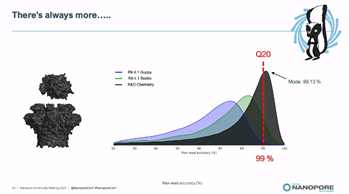
SR: This week Oxford Nanopore has generated modal *raw-read accuracy* of 99.1% (99%=Q20) using a new chemistry with Bonito, delivered on internal validation sets with a substantial fraction of these raw reads above Q20 #nanoporeconf

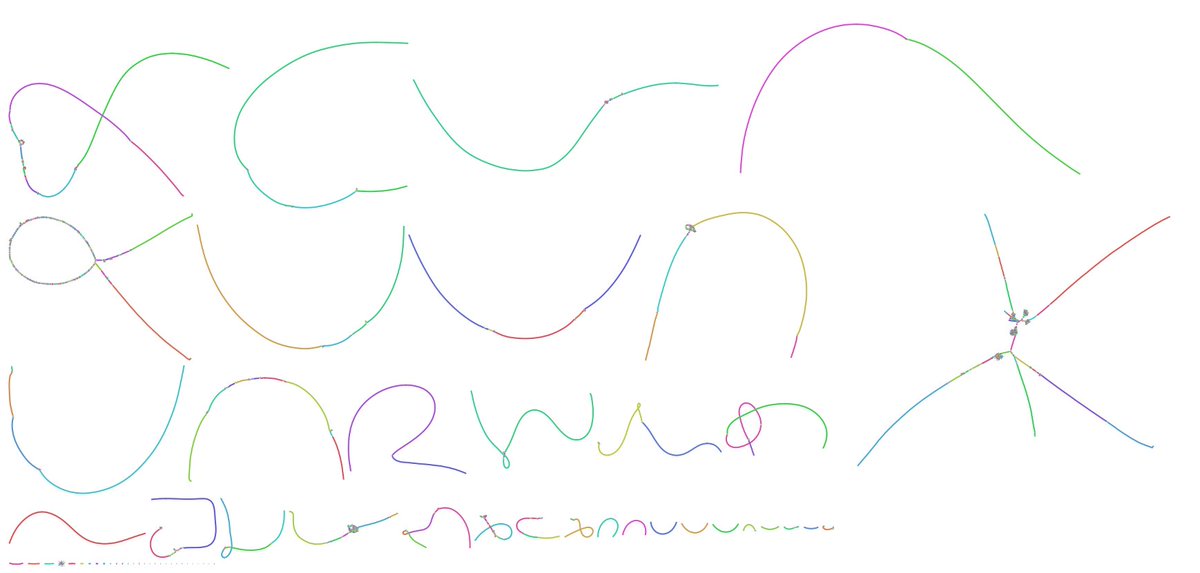
The Telomere-to-Telomere consortium is proud to announce our v1.0 release of a complete human genome. When Karen Miga asked me to team up on this a few years ago, I wasn't sure it would be possible. Now down to just 5 gaps. What a journey! Read more here genomeinformatics.github.io/CHM13v1/


Hifiasm, a haplotype-aware #HiFiReads assembler with builtin haplotig purging & much improved trio binning. Applied to human, mouse, maize, frog, octoploid strawberry & ~30Gb hexaploid redwood. By Haoyu Cheng, gconcepcion.bsky.social, Xiaowen Feng & Haowen Zhang at arxiv.org/abs/2008.01237

