
#TeamMassSpec are you ready for #ASMS2024 ? From incredible scientific content, technological breakthroughs, to unparalleled networking opportunities, see the full line-up of Thermo Fisher activities thermofisher.com/ASMS

Hi #TeamMassSpec #Proteomics
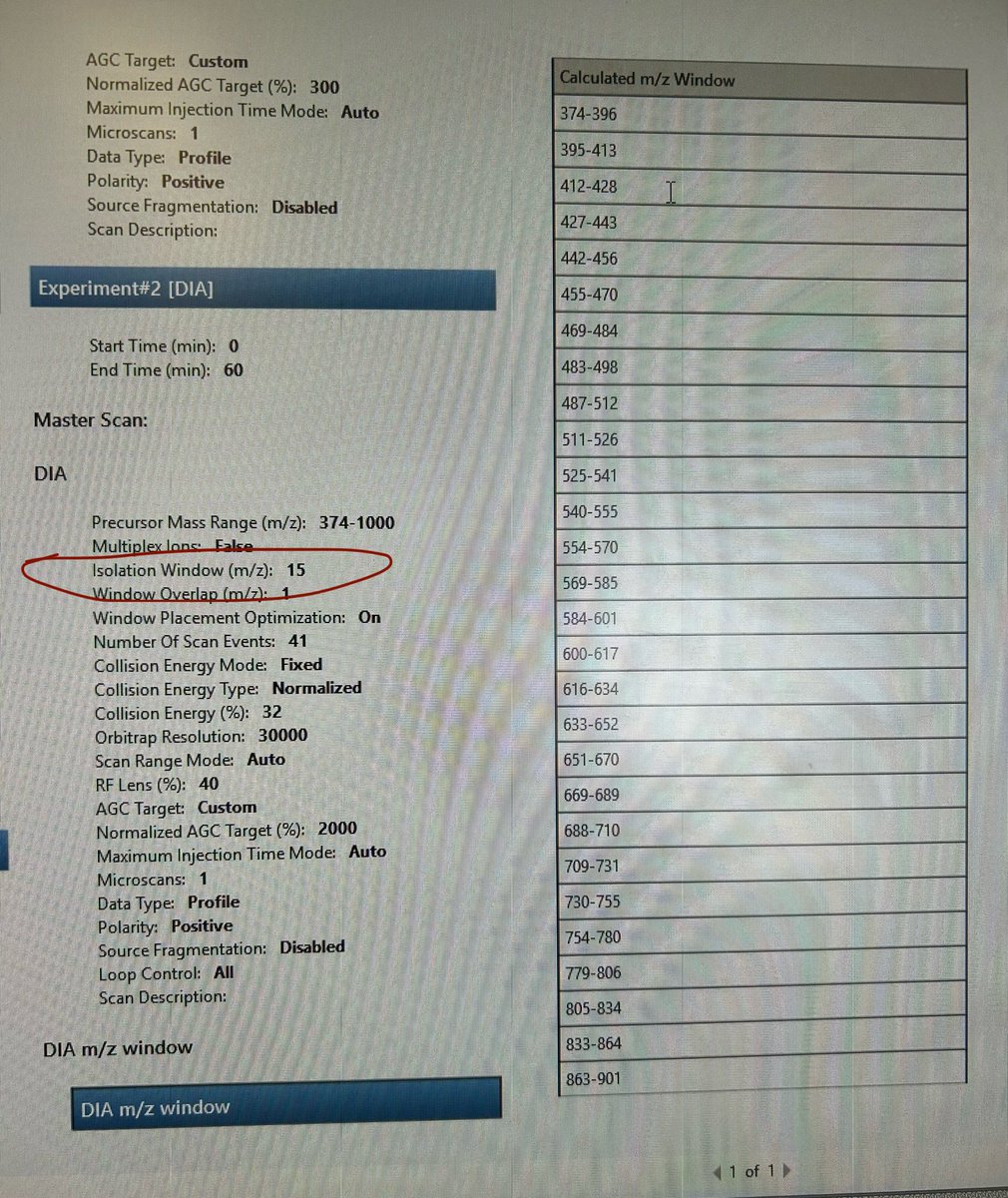
I'm attempting to create a variable m/z window (ranges from 14 to 56 m/z) DIA method on the Exploris 480. Is there a right approach to optimize the Isolation Window parameter from fixed (here it is 15) to variable?





Excited to give my talk at Am Soc for Mass Spec: “Deep Learning Enables Targeted Proteomics With Sample Multiplexing Without PRM Scheduling, Synthetic Peptides, or Data Libraries” Monday 6/3/24 at 3:50pm! Hope to see you there #teamMassSpec !


Brian Searle & Lindsay K Pino - @[email protected] presented a session at #ABRF2024 on 'Embracing Disruption & New Opportunities in the Evolution of #Proteomics Technologies' wherein #TeamMassSpec and Non-MassSpec applications were discussed. Slides will be available on the ABRF sPRG website soon.


Come and visit our next #AdvancedMassSpec seminar this Wednesday. The stage is ready for two outstanding talks in #proteomics & #metabolomics .
#TeamMassSpec #Bioinformatics


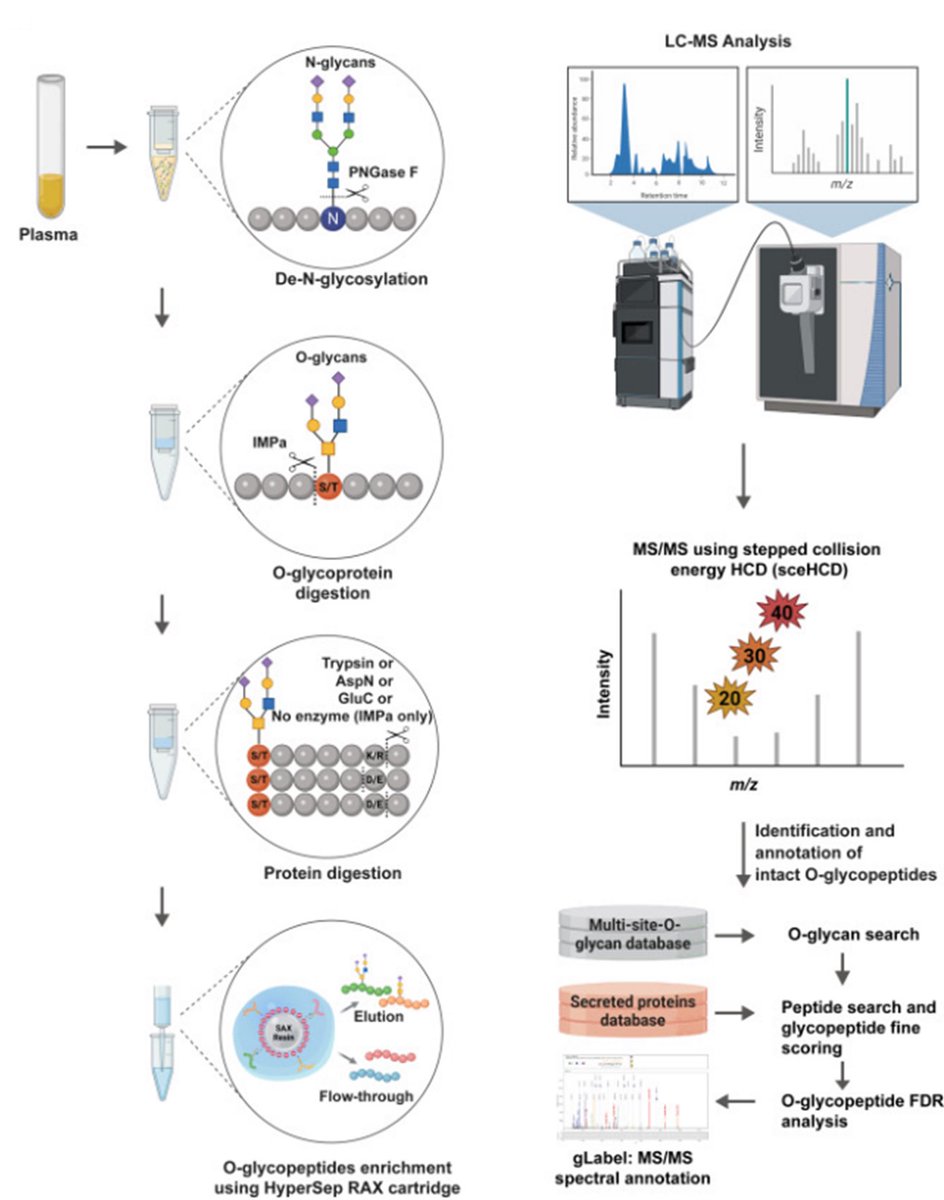
Excited about our recent paper in Cell Reports Methods on LC-MS/MS based global O-glycoproteome analysis in blood plasma co-led by Taewook Kang and Rohit Budhraja from our team #teammassspec #glycotime Cell Reports Methods Taewook Rohit budhraja

New Malaker lab photo just dropped! Welcome new grad students Ryan, Shane, and Ellie (Ellie Browne), as well as new undergrads Iris and George! 🥳🎉 #mucinome #TeamMassSpec #glycotime
Second photo is what we're really like...
~Photo cred to NaOH Bartfield 🧪~


In this video, discover how the HiLIFE Proteomics Unit at the University of Helsinki benefits from Evosep's dedicated service and support. Hear directly from PhD, Head of Laboratory Salla Keskitalo and her group as they detail their experience with our expert support team. #TeamMassSpec

Proteonano coupled with Orbitrap Astral: A robust and standardized workflow for large-cohort clinical plasma proteomics studies biorxiv.org/content/10.110…
A competition to SEER and possible Thermo acquisition 🤔 #Teammassspec

It was super fun to have Sharon Pitteri visit UW-Madison Chemistry today, delivering an amazing seminar on glycoproteomics for novel cancer diagnostics. We enjoyed the interactions and stimulating discussions. Lingjun Li Research Group Ying Ge #TeamMassSpec #Glycotime


Huge thanks to the teams from MOBILion Systems and Agilent Technologies who were on-site at IGFS this week to install and conduct initial training on the MOBIE, thanks to their experience, we unboxed the MOBIE on Tuesday and acquiring our first IM-MS data this afternoon. #TeamMassSpec



Calling the #singlecellproteomics #scp #teammassspec community to do some SCP analyses on CSM - key to better understand multicellularity, differentiation and systems biology fundamentals. Now we have the tools!
See video to spark your curiosity ⬇️
youtu.be/5h8WOWEqP6o?si…

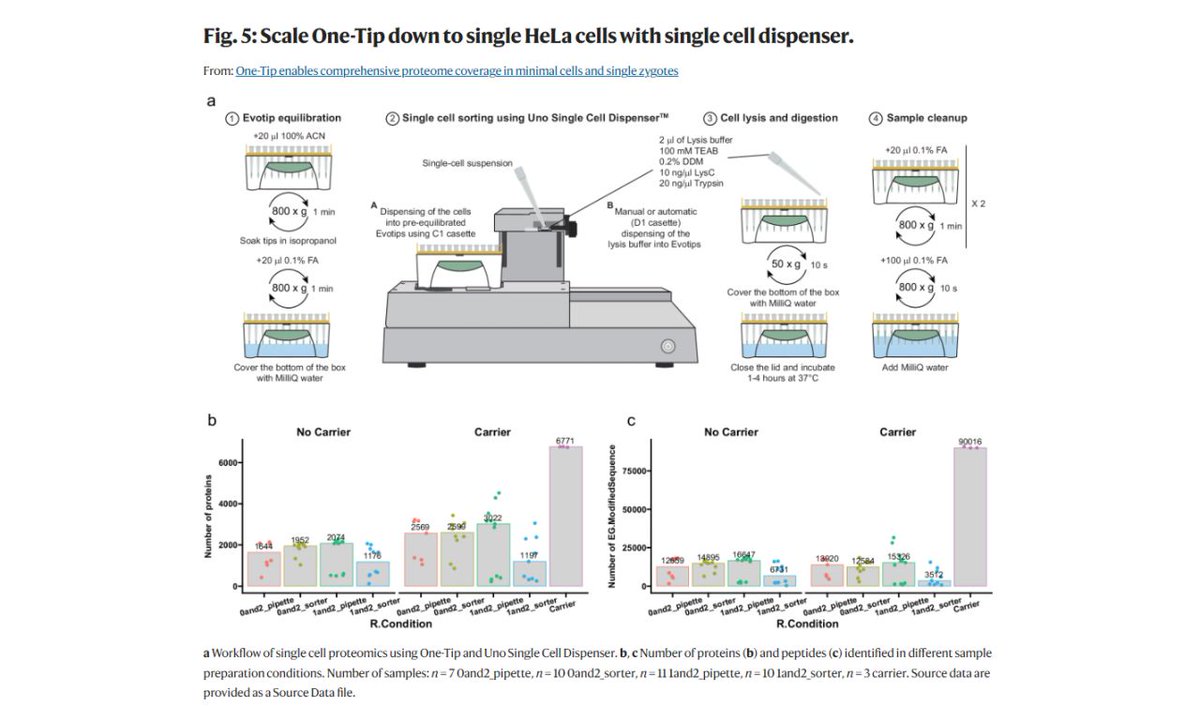
Visit or re-visit this established workflow with the One-Tip and how it gives you comprehensive #proteome coverage in minimal cells and single zygotes. nature.com/articles/s4146… #teammassspec #singlecellanalysis

Super excited for Matt, who will be defending his PhD this summer, for his selection for the NRC Research Associateship Programs to work with Patrick Fedick during his postdoc!
#TeamMassSpec #ProudPI Chemistry @ MSState

Latest #PubChemLite version out now!
zenodo.org/records/110702…
#cheminformatics #exposome #TeamMassSpec
Plus for CCS enthusiasts we're working with CCSbase UW XuLab and PubChem to get monthly PCL+predicted CCS running too (stay tuned, still WiP)!
zenodo.org/records/110578…



