
jdavieslab
@jdavieslab
Davies lab @MRC_WIMM @UniofOxford
ID:1309589916528062466
25-09-2020 20:26:11
92 Tweets
383 Followers
246 Following
Follow People


Very exited to share that our work showing the importance of SOX2 pioneer activity for gene activation has now been published in The EMBO Journal. Excellent work from MichEla and Teun van den Brand. And a nice collaboration with James Davies (+lab). embopress.org/doi/full/10.15…

Very happy and honored to present our work from my postdoc project in Noordermeer lab 🇺🇦 lab I2BC Paris-Saclay on CTCF clustering at TAD boundaries and the novel multi-contact 3C assay: Nano-C – now published Nature Communications.
nature.com/articles/s4146….
A nice 🧵 written by Daan 👇


We are excited to share our recent work mapping the host response to sepsis at single-cell resolution. The study, led by Andrew Kwok and published in Nature Immunology, identifies a subset of immature neutrophils which is increased in severe sepsis.
Big thanks to everyone involved!

The Clinicogenomic Landscape of Induction Failure in Childhood and Young Adult T-Cell Acute Lymphoblastic Leukemia ascopubs.org/doi/10.1200/JC…
TAL1 non coding mutations predict induction failure and poor outcome in T-ALL.
Terrific work from Marc Mansour and colleagues.

#efficacy #genome #editing exa-cel Vertex Pharmaceuticals leading to abrogation disease manifestations most #patients #thalassaemia & #sicklecell #EBMT2023 The EBMT #presidential ; Imperial NHS 💙 Imperial Department of Immunology & Inflammation part of this work
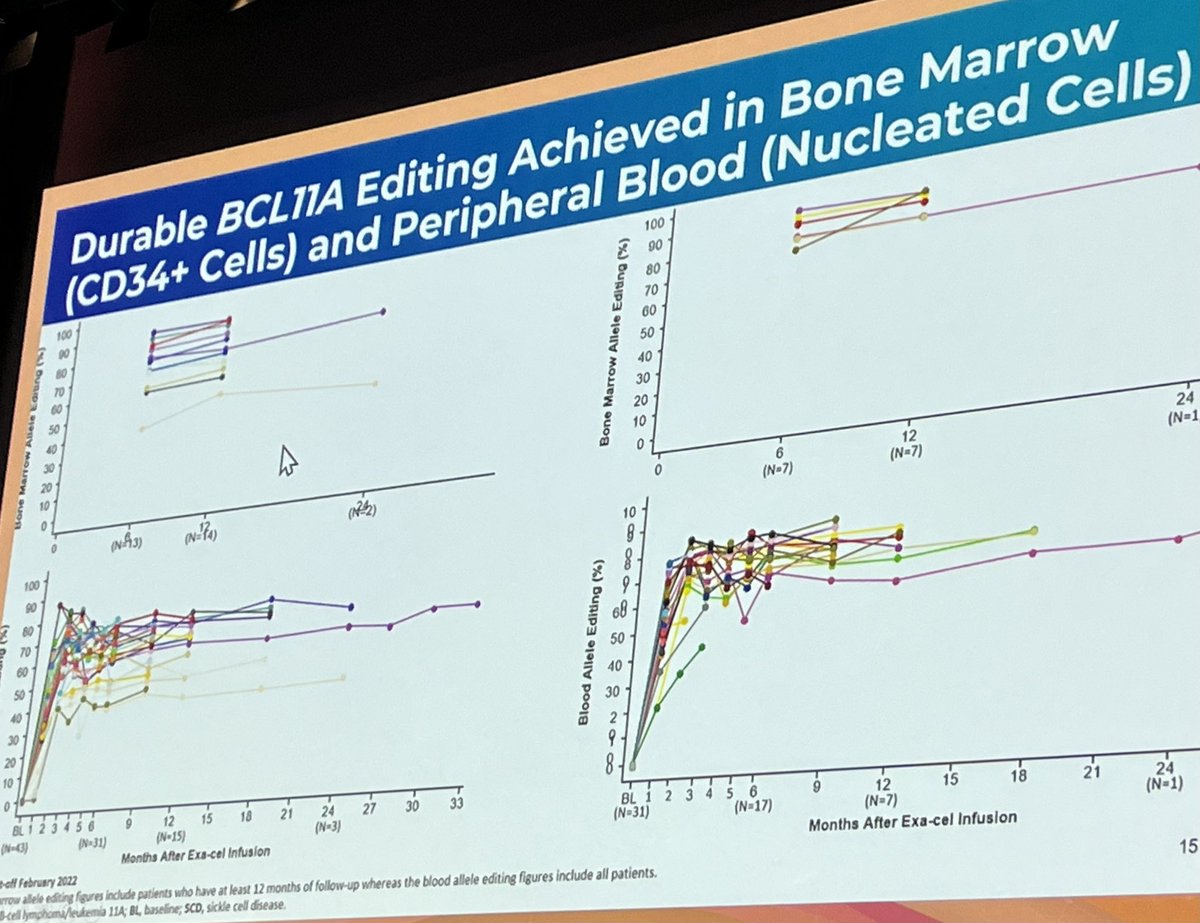


In 2016 I looked after a girl who died during a transplant for HbE beta-thalassaemia.
I am overwhelmingly pleased that we have been able to correct HbE with near 100% efficiency using base editors.
nature.com/articles/s4146…
Brilliant work from Mohsin Badat Mohsin Badat jdavieslab

The Huntly lab CRUK Cambridge Centre: Haematological Malignancies has an opening for a research associate in bioinformatics. Great science, tonnes of data and a lovely team. Please RT.
jobs.cam.ac.uk/job/40423/

Really pleased to see our Nature Protocols paper out this week. We have put together a comprehensive guide to generating base resolution 3C data!
Hangpeng Li James Davies jdavieslab

Congratulations Joseph Hamley and Hangpeng Li for putting together a nature protocols paper for Micro Capture-C! nature.com/articles/s4159…

Cofactor requirements for transcription factors! A labour of love for Charlie Bell - so very proud of him and all in The Dawson Lab involved. A body of work that is not only amazing in scale but also tackles an important and tough question in our field - Enjoy!



Only ~2% of the human genome codes for proteins.
A Wellcome Discovery award for Jim R. Hughes jdavieslab and Cecilia Lindgren aims to deepen our understanding of the other 98% and its role in disease.
Big Data Institute MRC WIMM
Music: MNDLSS

A new Wellcome Discovery award project aims to use Micro Capture-C and Deep Neural Network Machine Learning to decipher the role of the non-coding genome in disease.
Jim R. Hughes. jdavieslab MRC MHU
Cecilia Lindgren Big Data Institute
bit.ly/3D7B1oK

Really excited to work on this project with Cecilia Lindgren and Jim R. Hughes! Wellcome wellcome.org/grant-funding/…






